Exploratory Data Analysis
BIOSTAT 620: Introduction to Health Data Science
Exploratory Data Analysis
Main topic for today
Acknowledgment
This lecture includes some slides originally developed by Meredith Franklin. They have been modified by George G. Vega Yon, Kelly Street, and Dylan Cable.
Exploratory Data Analysis
- Exploratory data analysis is the process of becoming familiar with a dataset
- It should be the first step in your analysis pipeline
- It involves:
- checking data (import issues, outliers, missing values, data errors)
- cleaning data
- summary statistics of key variables (univariate and bivariate)
- basic plots and graphs
Exploratory Data Analysis
Since our eyes and brains are not wired to detect patterns in large data tables filled with text and numbers, communication about data […] rarely comes in the form of raw data or code output. Instead, data and data-driven results are usually either summarized (e.g., using an average/mean) and presented in small summary tables or they are presented visually in the form of graphs, in which shape, distance, color, and size can be used to represent the magnitudes of (and relationships between) the values within our data.
- Viridical Data Science, Yu and Barter
Key Questions
- What question(s) am I trying to answer?
- What question(s) could this dataset answer?
The Tidyverse Model
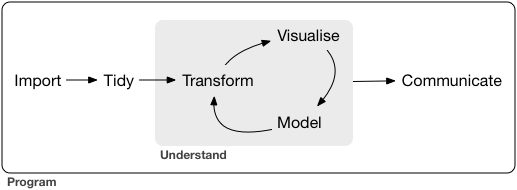
Loosely, EDA encompasses the Import -> Tidy -> Transform -> Visualize steps. Basically it is everything before we do modeling, prediction or inference.
EDA may involve some statistical summaries, but it does not include formal statistical analysis.
EDA Checklist
The goal of EDA is to better understand your data. Let’s use the checklist:
- Read in the data
- Check the size of the data
- Examine the variables and their types
- Look at the top and bottom of the data
- Visualize the distributions of key variables
- Check your expectations
- Validate with an external source
- Formulate a (simple) question
- Try the easy solution first
- Challenge your solution
(adapted from Exploratory Data Analysis with R by Roger D. Peng)
Case study
We are going to use a dataset created from the National Center for Environmental Information (https://www.ncei.noaa.gov/). The data are 2019 hourly measurements from weather stations across the continental U.S.
Formulate a Question
It is a good idea to first have a question such as:
- Which weather stations reported the hottest and coldest daily temperatures?
- What day of the month was on average the hottest?
- Is there correlation between temperature and humidity in my dataset?
Read in the Data
There are several ways to read in data (some depend on the type of data you have):
read.tableorread.csvin base R for delimited filesreadRDSif you have a .rds dataset (this is a handy, compressed way of saving R objects)read_csv,read_csv2,read_delim,read_fwffromlibrary(readr)that is part of the tidyversereadxl()fromlibrary(readxl)for .xls and .xlsx filesread_sas,read_spss,read_statafromlibrary(haven)freadfromlibrary(data.table)for efficiently importing large datasets that are regular delimited files
Read in the Data
There are plenty of ways to do these tasks, but we will focus on base R.
Since our data is stored as a (gzipped) CSV file, we could load it into R with read.csv, but we will use the more flexible read.table. I have it stored locally, but we will see how to load it straight from GitHub in the lab.
We specify that the first line contains column names by setting header = TRUE and we indicate that commas are used to separate the different values (rather than tabs, spaces, etc.) by setting sep = ','.
Working with data.frames
This gave as a data.frame object, which is a standard R format for cleaned, rectangular data. Each row represents an observation and each column represents a variable.
As we have seen, you can access particular parts of the data.frame by subsetting with the square brackets, [,]. For example, you can pull out the 2nd, 3rd, and 4th elements of the 1st column of our met dataset with met[2:4, 1].
You can also pull out specific columns by name, using the $ operator. Since the first column is called USAFID, we could access the same subset as above with met$USAFID[2:4] (notice that there is no comma here, because we have already subset down to a single variable).
To see the list of names for the dataset, you can use names(met) or colnames(met). To see the top few rows of the dataset, use head(met).
Check the data
We should check the dimensions of the data set. This can be done several ways:
Check the data
- We see that there are 2,377,343 records of hourly temperature in August 2019 from all of the weather stations in the US. The data set has 30 variables.
- We should also check the top and bottom of the dataset to check for any irregularities. Use
head(met)andtail(met)for this. - Next we can take a deeper dive into the contents of the data with
str()
Check variables
'data.frame': 2377343 obs. of 30 variables:
$ USAFID : int 690150 690150 690150 690150 690150 690150 690150 690150 690150 690150 ...
$ WBAN : int 93121 93121 93121 93121 93121 93121 93121 93121 93121 93121 ...
$ year : int 2019 2019 2019 2019 2019 2019 2019 2019 2019 2019 ...
$ month : int 8 8 8 8 8 8 8 8 8 8 ...
$ day : int 1 1 1 1 1 1 1 1 1 1 ...
$ hour : int 0 1 2 3 4 5 6 7 8 9 ...
$ min : int 56 56 56 56 56 56 56 56 56 56 ...
$ lat : num 34.3 34.3 34.3 34.3 34.3 34.3 34.3 34.3 34.3 34.3 ...
$ lon : num -116 -116 -116 -116 -116 ...
$ elev : int 696 696 696 696 696 696 696 696 696 696 ...
$ wind.dir : int 220 230 230 210 120 NA 320 10 320 350 ...
$ wind.dir.qc : chr "5" "5" "5" "5" ...
$ wind.type.code : chr "N" "N" "N" "N" ...
$ wind.sp : num 5.7 8.2 6.7 5.1 2.1 0 1.5 2.1 2.6 1.5 ...
$ wind.sp.qc : chr "5" "5" "5" "5" ...
$ ceiling.ht : int 22000 22000 22000 22000 22000 22000 22000 22000 22000 22000 ...
$ ceiling.ht.qc : int 5 5 5 5 5 5 5 5 5 5 ...
$ ceiling.ht.method: chr "9" "9" "9" "9" ...
$ sky.cond : chr "N" "N" "N" "N" ...
$ vis.dist : int 16093 16093 16093 16093 16093 16093 16093 16093 16093 16093 ...
$ vis.dist.qc : chr "5" "5" "5" "5" ...
$ vis.var : chr "N" "N" "N" "N" ...
$ vis.var.qc : chr "5" "5" "5" "5" ...
$ temp : num 37.2 35.6 34.4 33.3 32.8 31.1 29.4 28.9 27.2 26.7 ...
$ temp.qc : chr "5" "5" "5" "5" ...
$ dew.point : num 10.6 10.6 7.2 5 5 5.6 6.1 6.7 7.8 7.8 ...
$ dew.point.qc : chr "5" "5" "5" "5" ...
$ atm.press : num 1010 1010 1011 1012 1013 ...
$ atm.press.qc : int 5 5 5 5 5 5 5 5 5 5 ...
$ rh : num 19.9 21.8 18.5 16.9 17.4 ...Check variables
- First, we see that
str()gives us the class of the data, which in this case is adata.frame, as well as the dimensions of the data - We also see the variable names and their type (integer, numeric, character, etc.)
- We can identify major problems with the data at this stage (e.g. a variable that has all missing values)
Check variables
We can get summary statistics on our data.frame using summary().
lat lon elev wind.dir
Min. :24.55 Min. :-124.29 Min. : -13.0 Min. : 3
1st Qu.:33.97 1st Qu.: -98.02 1st Qu.: 101.0 1st Qu.:120
Median :38.35 Median : -91.71 Median : 252.0 Median :180
Mean :37.94 Mean : -92.15 Mean : 415.8 Mean :185
3rd Qu.:41.94 3rd Qu.: -82.99 3rd Qu.: 400.0 3rd Qu.:260
Max. :48.94 Max. : -68.31 Max. :9999.0 Max. :360
NA's :785290
wind.dir.qc wind.type.code
Length:2377343 Length:2377343
Class :character Class :character
Mode :character Mode :character
Check variables more closely
We know that we are supposed to have hourly measurements of weather data for the month of August 2019 for the entire US. We should check that we have all of these components. Let’s check:
- the year
- the month
- the hours
- the range of locations (latitude and longitude)
Check variables more closely
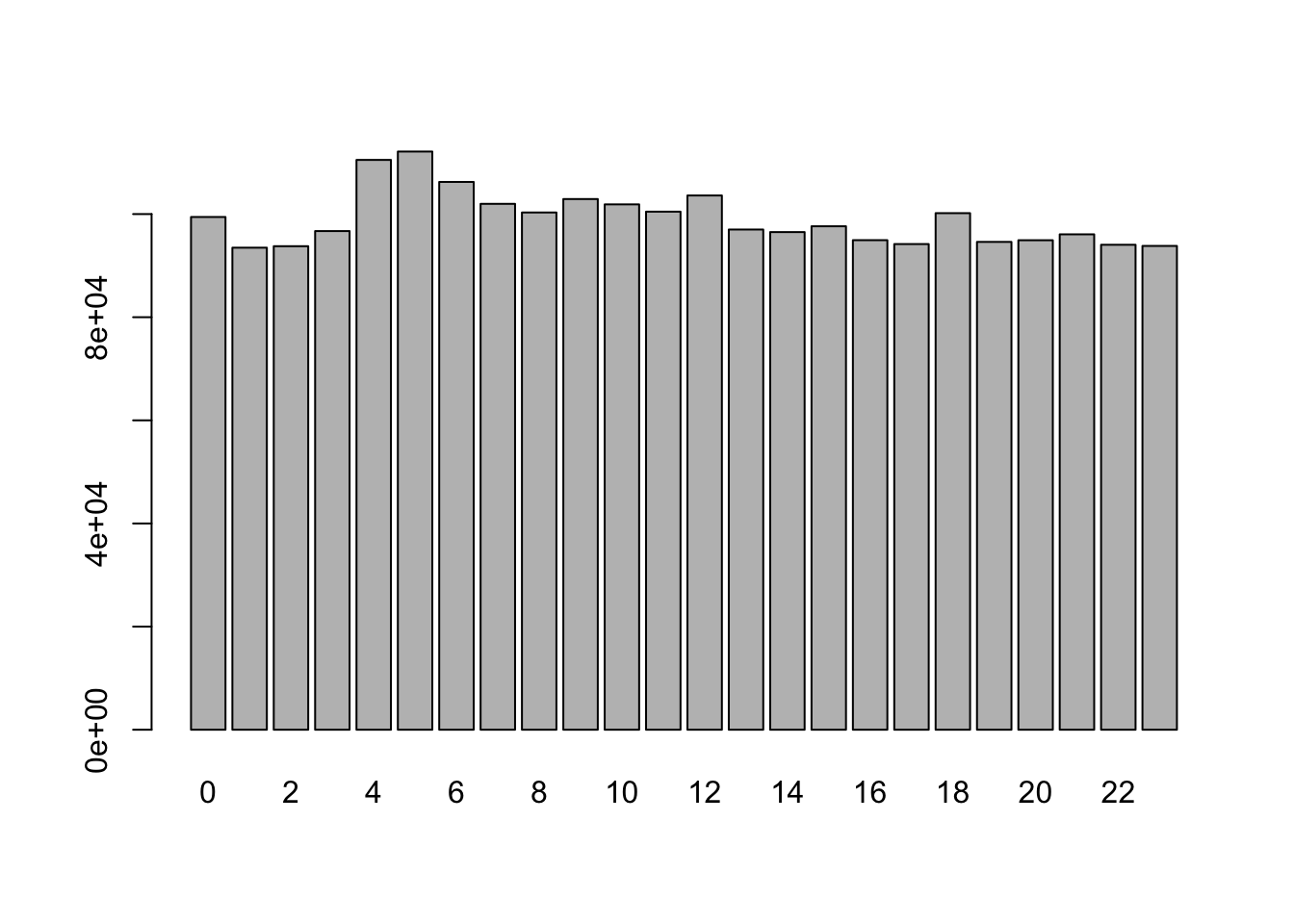
We can generate tables and/or barplots for integer variables:
Check variables more closely
We can generate tables and/or barplots for integer variables:
Check variables more closely
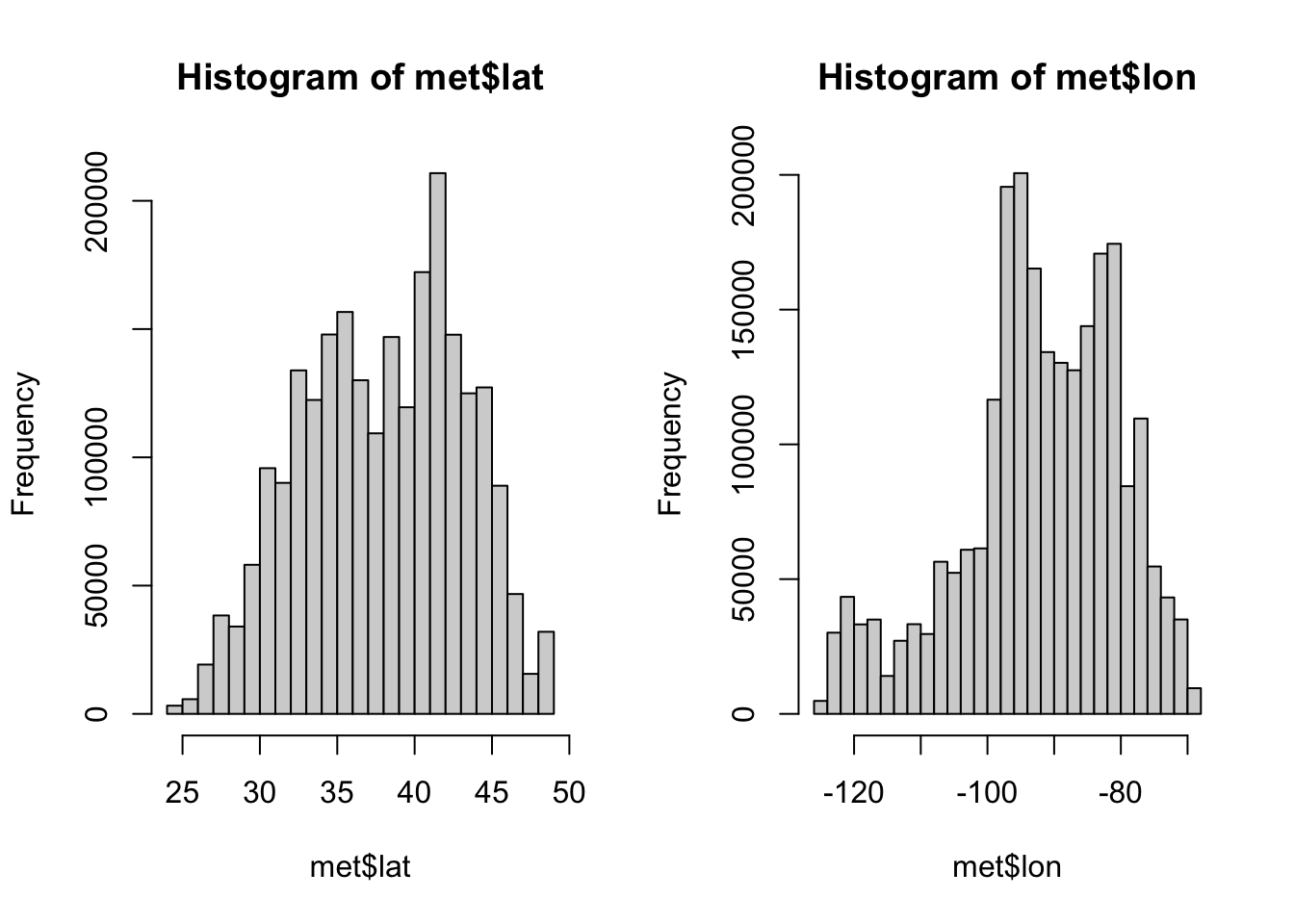
For numeric variables we should do a summary to see the quantiles, min, max, and mean.
Check variables more closely
We can visualize these distributions with a histogram.
Check variables more closely
If we return to our initial question, what weather stations reported the hottest and coldest temperatures, we should take a closer look at our key variable, temperature (temp)
Min. 1st Qu. Median Mean 3rd Qu. Max. NA's
-40.00 19.60 23.50 23.59 27.80 56.00 60089 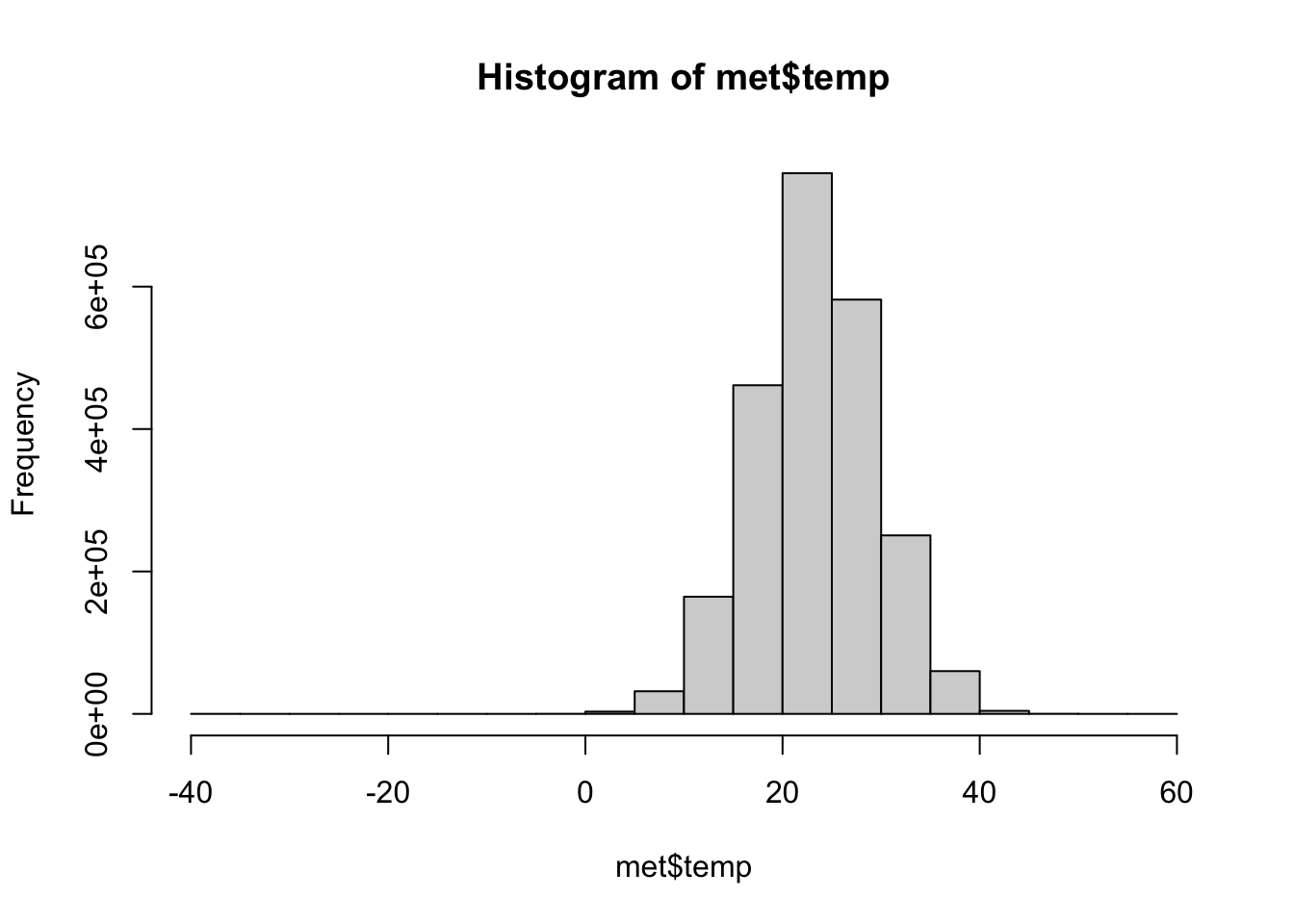
It looks like the temperatures are in Celsius. A temperature of -40 in August is really cold, we should see if this is an implausible value.
Check variables more closely
It also looks like there is a lot of missing data, encoded by NA values. Let’s check the proportion of missingness by tallying up whether or not every temperature reading is an NA. This will give us a vector of TRUE/FALSE values and then we can take the mean (average), because R automatically interprets TRUE as 1 and FALSE as 0 for mathematical functions.
2.5% of the data are missing, which is not a huge amount.
Check variables more closely
In our data.frame we can easily subset the data and select certain columns. Here, we select all observations with a temperature of -40C and a specific subset of the variables:
[1] 60125 5 hour lat lon elev
Min. : 0.00 Min. :29.12 Min. :-89.55 Min. :36
1st Qu.: 2.75 1st Qu.:29.12 1st Qu.:-89.55 1st Qu.:36
Median : 5.50 Median :29.12 Median :-89.55 Median :36
Mean : 5.50 Mean :29.12 Mean :-89.55 Mean :36
3rd Qu.: 8.25 3rd Qu.:29.12 3rd Qu.:-89.55 3rd Qu.:36
Max. :11.00 Max. :29.12 Max. :-89.55 Max. :36
NA's :60089 NA's :60089 NA's :60089 NA's :60089
wind.sp
Min. : NA
1st Qu.: NA
Median : NA
Mean :NaN
3rd Qu.: NA
Max. : NA
NA's :60125 Check variables more closely
In dplyr we can do the same thing using filter and select
library(dplyr)
met_ss <- filter(met, temp == -40.00) |>
select(USAFID, day, hour, lat, lon, elev, wind.sp)
dim(met_ss)[1] 36 7 USAFID day hour lat lon
Min. :720717 Min. :1 Min. : 0.00 Min. :29.12 Min. :-89.55
1st Qu.:720717 1st Qu.:1 1st Qu.: 2.75 1st Qu.:29.12 1st Qu.:-89.55
Median :720717 Median :1 Median : 5.50 Median :29.12 Median :-89.55
Mean :720717 Mean :1 Mean : 5.50 Mean :29.12 Mean :-89.55
3rd Qu.:720717 3rd Qu.:1 3rd Qu.: 8.25 3rd Qu.:29.12 3rd Qu.:-89.55
Max. :720717 Max. :1 Max. :11.00 Max. :29.12 Max. :-89.55
elev wind.sp
Min. :36 Min. : NA
1st Qu.:36 1st Qu.: NA
Median :36 Median : NA
Mean :36 Mean :NaN
3rd Qu.:36 3rd Qu.: NA
Max. :36 Max. : NA
NA's :36 Validate against an external source
We should check outside sources to make sure that our data makes sense. For example the observation with -40C is suspicious, so we should look up the location of the weather station.
Go to Google maps and enter the coordinates for the site with -40C (29.12, -89.55)
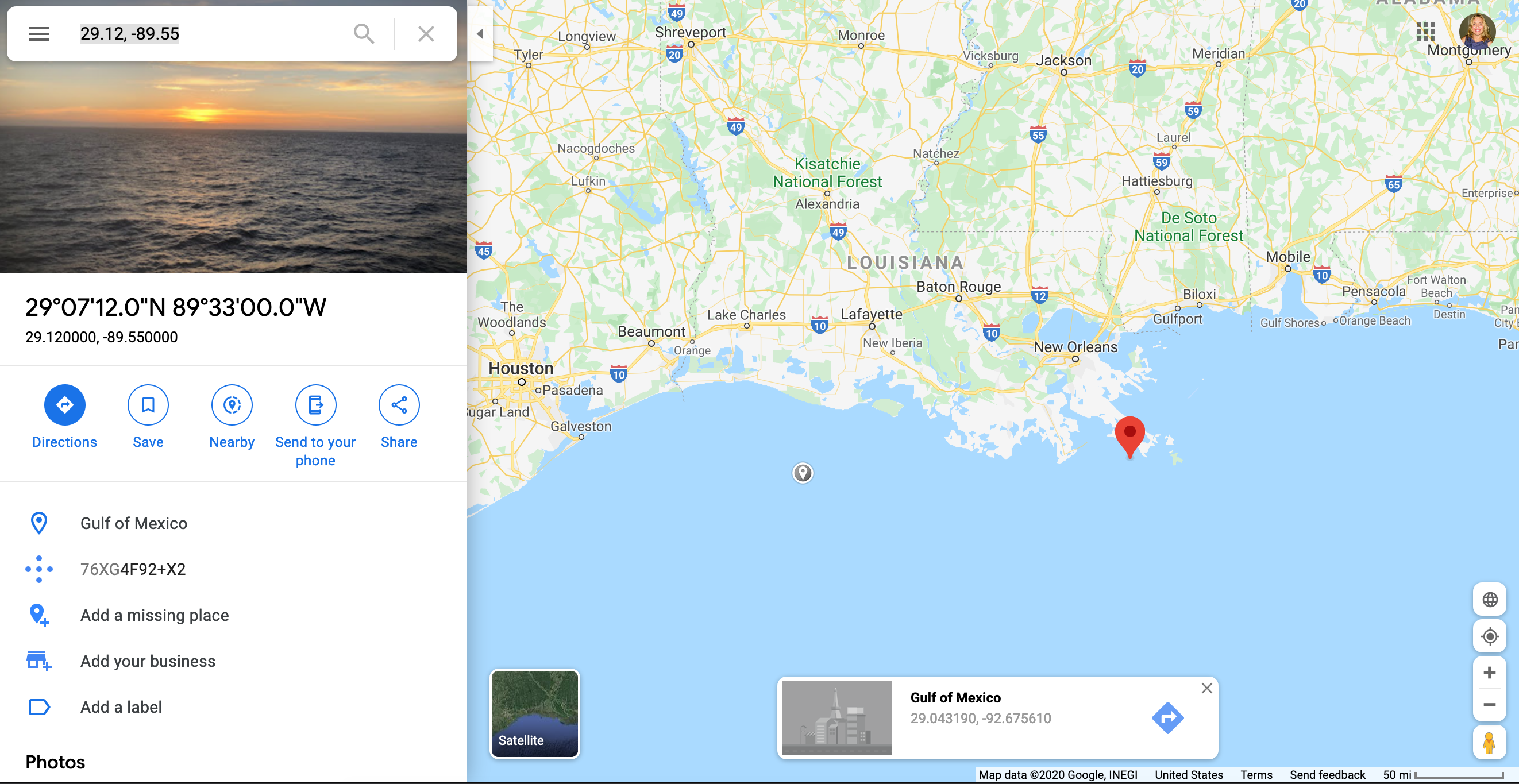
It doesn’t make much sense to have a -40C reading in the Gulf of Mexico off the coast of Louisiana!
Data cleaning
If we return to our initial question (“Which weather stations reported the hottest and coldest daily temperatures?”), we need to generate a list of weather stations that are ordered from highest to lowest. We can then examine the top and bottom of this new dataset.
First let us remove the aberrant observations and then we’ll sort by temperature.
Notice that we do not create a new object, we just overwrite the met object. Once you’re sure that you want to remove certain observations, this is a good way to avoid confusion (otherwise, it is easy to end up with multiple subsets of the data in your R environment with similar names like met, met_ss, met_ss2, met_final, met_FINAL, met_FINAL_REAL, etc.)
Data cleaning
We will also remove any observations with missing temperature values (NA).
The is.na() function tells you whether or not a particular value is missing and the ! operator takes the opposite of a TRUE/FALSE value, so in combination, they tell you which observations are not missing.
Sorting
Now, we can use the order() function to sort our dataset.
Again, we just replace the met object with this updated version, since we aren’t actually losing any data, just changing the order.
Highest and Lowest
USAFID lat lon elev temp
1203053 722817 38.767 -104.3 1838 -17.2
1203055 722817 38.767 -104.3 1838 -17.2
1203128 722817 38.767 -104.3 1838 -17.2
1203129 722817 38.767 -104.3 1838 -17.2
1203222 722817 38.767 -104.3 1838 -17.2
1203225 722817 38.767 -104.3 1838 -17.2 USAFID lat lon elev temp
42783 720267 38.955 -121.081 467 52.0
724 690150 34.300 -116.166 696 52.8
749 690150 34.296 -116.162 625 52.8
748 690150 34.300 -116.166 696 53.9
701 690150 34.300 -116.166 696 54.4
42403 720267 38.955 -121.081 467 56.0Summary statistics
The maximum hourly temperature is 56C at site 720267, and the minimum hourly temperature is -17.2C at site 722817.
Summary statistics
We need to transform our data to answer our initial question. Let’s find the daily mean, max, and min temperatures for each weather station in our data.frame. We can do this with the summarize function from the dplyr package. This package is part of the tidyverse, so the syntax is a bit different from what we’ve seen before.
What we’ve done here is told R to summarize the met dataset by the variables USAFID and day, splitting the data into subsets based on those two indexing variables. For each subset (representing a specific station of a specific day), we want the daily average temperature, as well as latitude, longitude, and elevation (though hopefully those don’t change too much over the course of a day!)
Summary statistics
Before we continue, check the relative sizes of the met and met_daily objects. Which one is bigger?
Summary statistics
Now we will order our new dataset by the average daily temperature, just as we ordered the old one by observed temperature.
USAFID day temp lat lon elev
2 722817 3 -17.200000 38.767 -104.3 1838
1 722817 1 -17.133333 38.767 -104.3 1838
3 722817 6 -17.066667 38.767 -104.3 1838
164 726130 11 4.278261 44.270 -71.3 1909
166 726130 31 4.304348 44.270 -71.3 1909
163 726130 10 4.583333 44.270 -71.3 1909 USAFID day temp lat lon elev
48708 722749 5 40.85714 33.26900 -111.8120 379.0000
48695 723805 5 40.97500 34.76800 -114.6180 279.0000
48721 720339 14 41.00000 32.14600 -111.1710 737.0000
48710 723805 4 41.18333 34.76800 -114.6180 279.0000
48688 722787 5 41.35714 33.52700 -112.2950 325.0000
48438 690150 31 41.71667 34.29967 -116.1657 690.0833Summary statistics
The maximum daily average temperature is 41.7166667 C at site 690150 and the minimum daily average temperature is -17.2C at site 722817.
Summary statistics
The code below is similar to our previous example, but doesn’t include the latitude, longitude, and elevation. How would you alter this code to find the daily median, max, or min temperatures for each station?
(try it yourself)
Exploratory graphs
With exploratory graphs we aim to:
- debug any issues remaining in the data
- understand properties of the data
- look for patterns in the data
- inform modeling strategies
Exploratory graphs do not need to be perfect, we will look at presentation ready plots next week.
Exploratory graphs
Examples of exploratory graphs include:
- histograms
- boxplots
- scatterplots
- simple maps
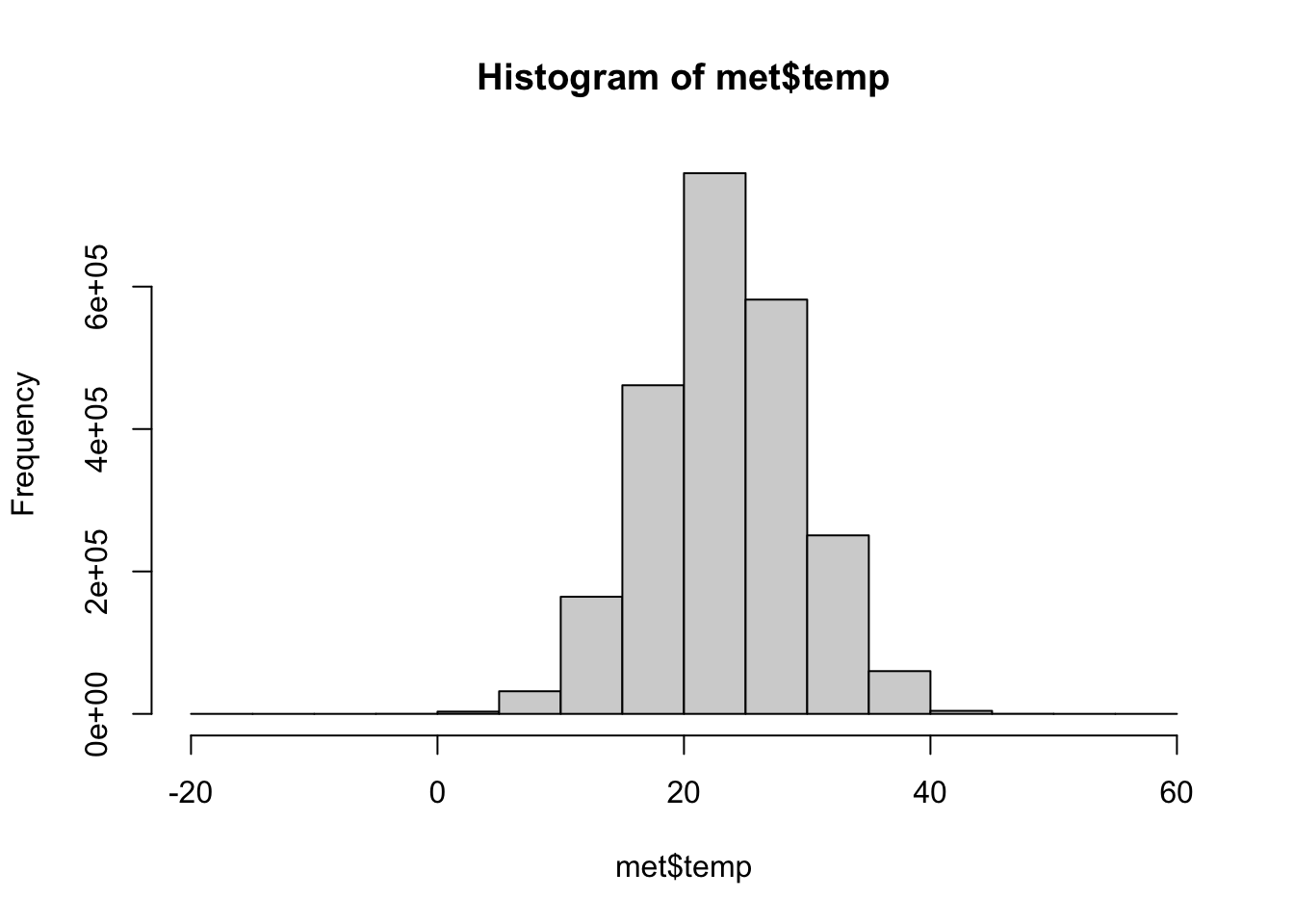
Exploratory Graphs
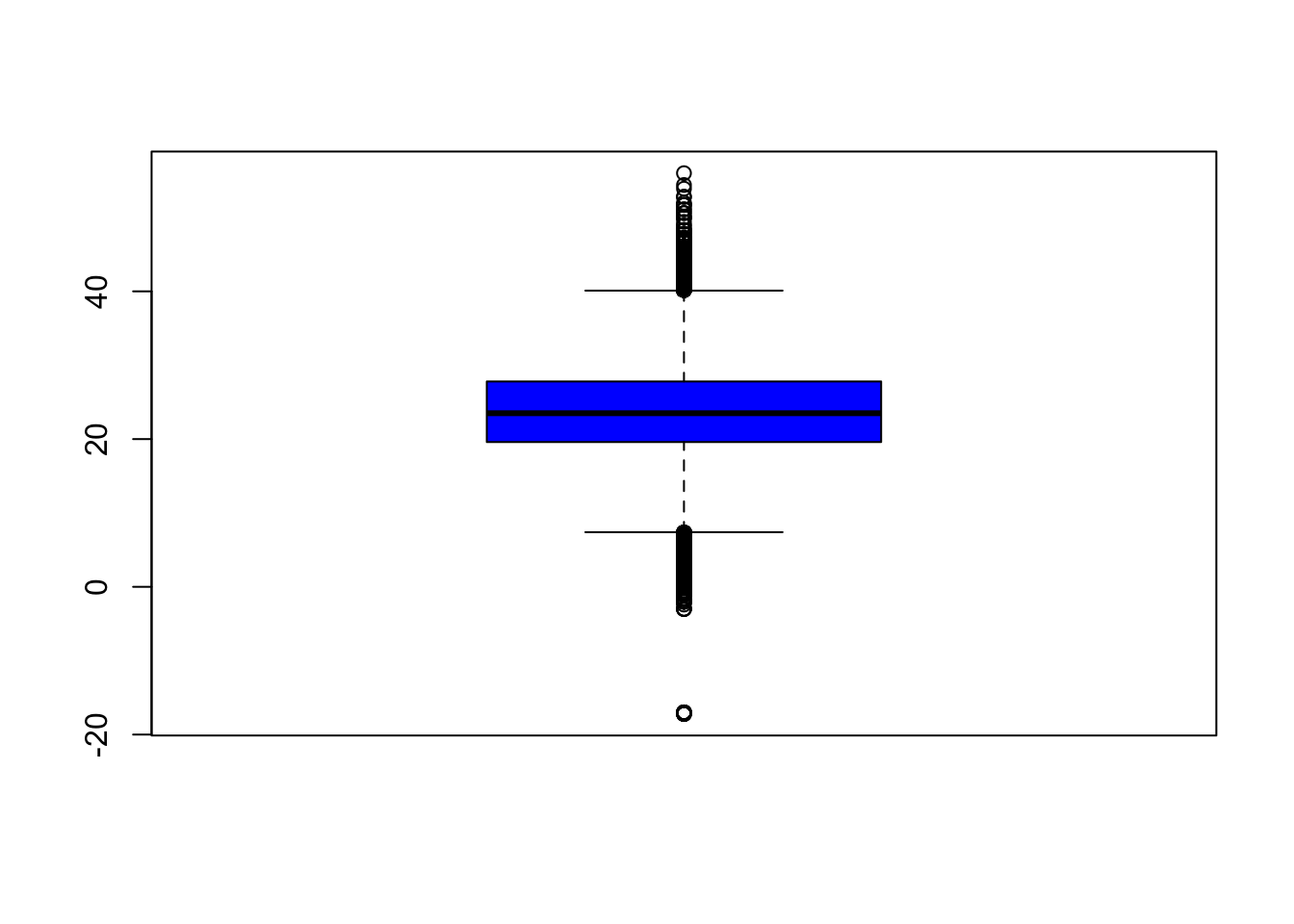
Focusing on the variable of interest, temperature, let’s look at the distribution (after removing -40C)
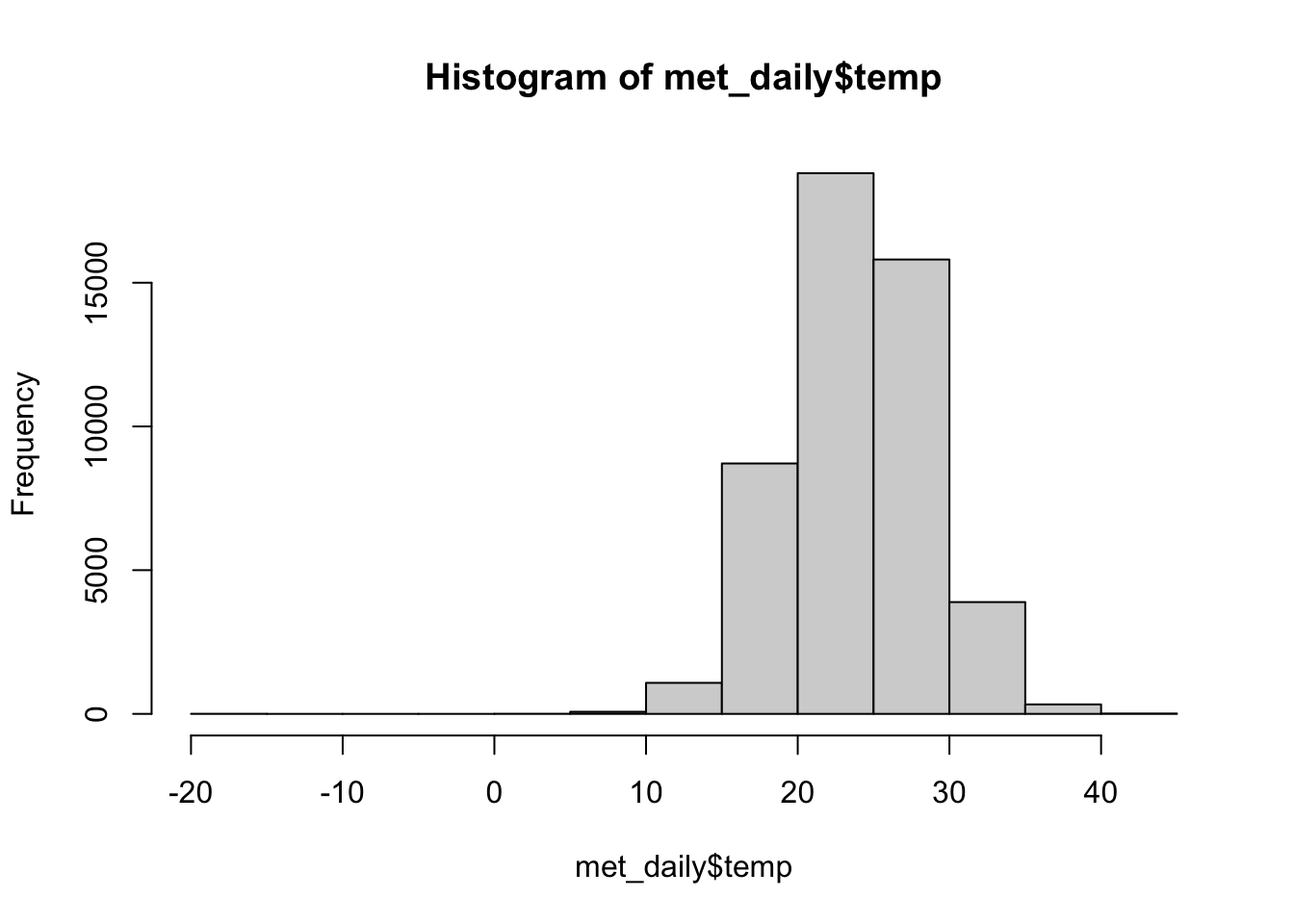
Exploratory Graphs
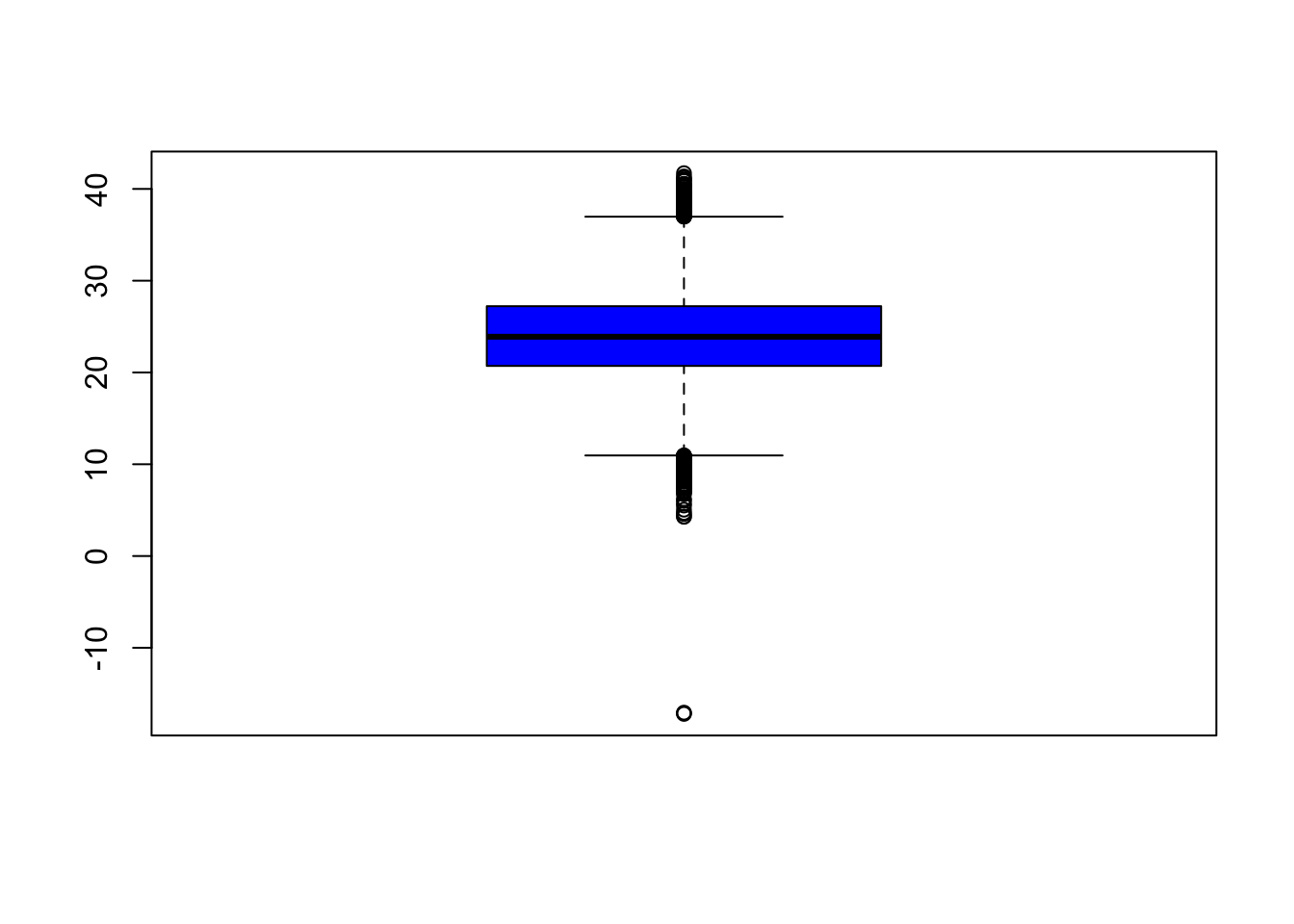
Let’s look at the daily data
Exploratory Graphs
A boxplot gives us an idea of the quantiles of the distribution and any outliers
Exploratory Graphs
Let’s look at the daily data
Exploratory Graphs
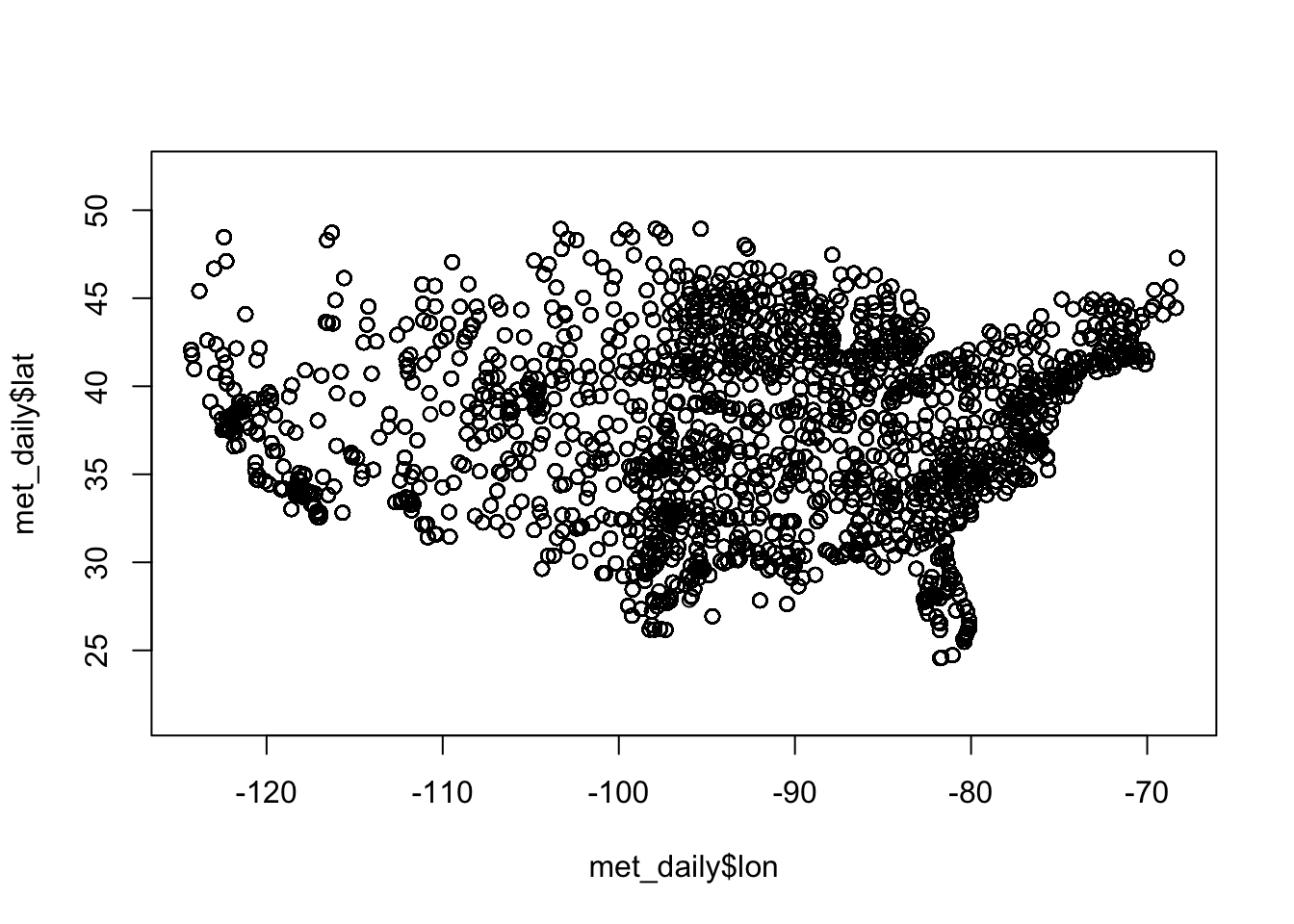
We know that these data come from US weather stations, so we might have some idea what to expect just from plotting the latitude and longitude (note that we fix the aspect ratio at 1:1 with asp = 1; this prevents the plot from stretching or shrinking to fit the available plotting area):
Exploratory Graphs
A map will show us where the weather stations are located. First let’s get the unique latitudes and longitudes and see how many meteorological sites there are
Exploratory Graphs
A map will show us where the weather stations are located. First let’s get the unique latitudes and longitudes and see how many meteorological sites there are.
Exploratory Graphs
Exploratory Graphs
Let’s map the locations of the max and min daily temperatures.
min <- met_daily[1, ] # First observation
max <- met_daily[nrow(met_daily), ] # Last observation
leaflet() |>
addProviderTiles('CartoDB.Positron') |>
addCircles(
data = min,
lat = ~lat, lng = ~lon, popup = "Min temp.",
opacity = 1, fillOpacity = 1, radius = 400, color = "blue"
) |>
addCircles(
data = max,
lat = ~lat, lng = ~lon, popup = "Max temp.",
opacity=1, fillOpacity=1, radius = 400, color = "red"
)(next slide)
Exploratory Graphs
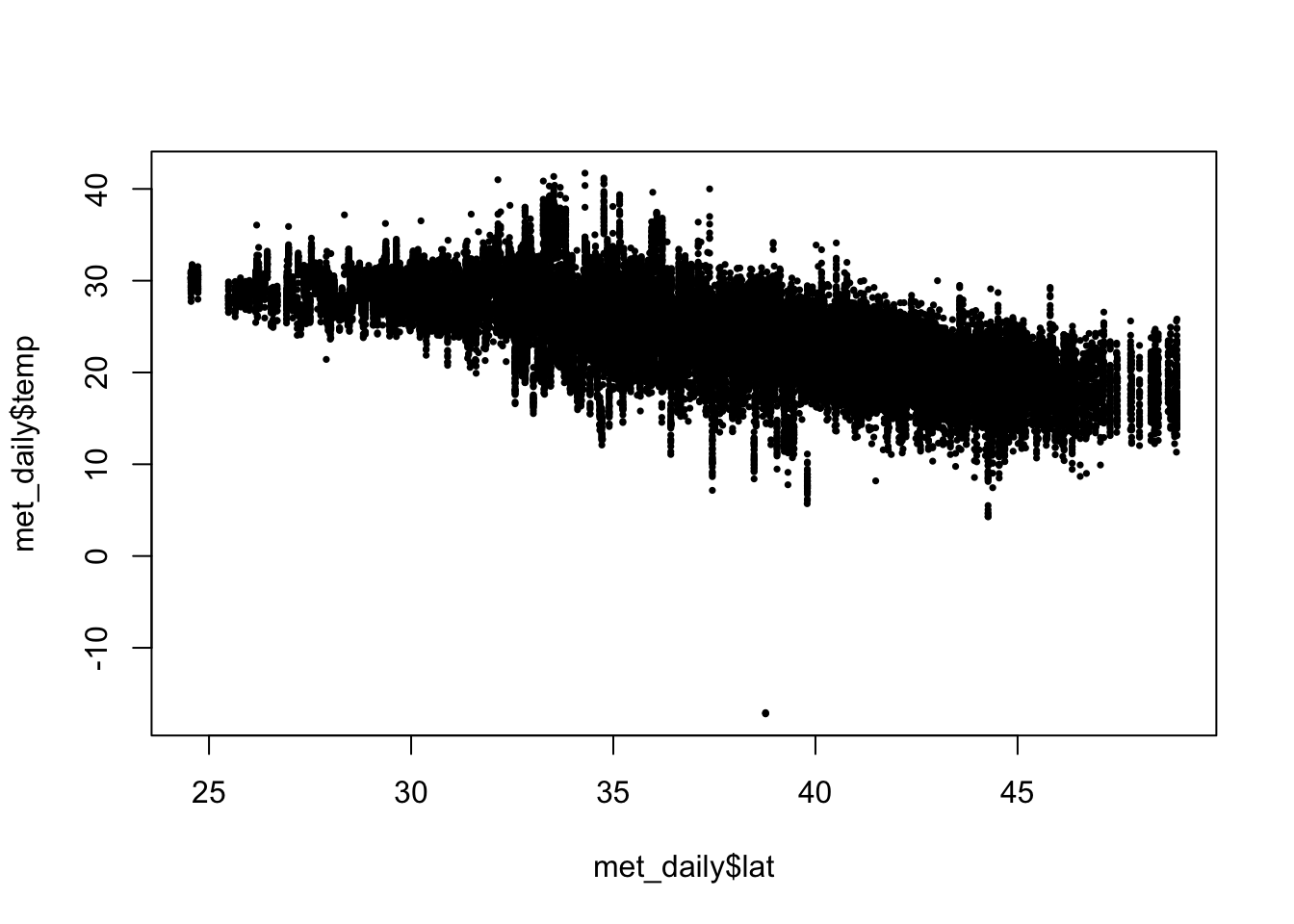
Scatterplots help us look at pairwise relationships. Let’s see if there is any trend in temperature with latitude
There is a clear decrease in temperatures as you increase in latitude (i.e as you go north).
Exploratory Graphs
We can add a simple linear regression line to this plot using lm() and abline(). We can also add a title and change the axis labels.
(next slide)
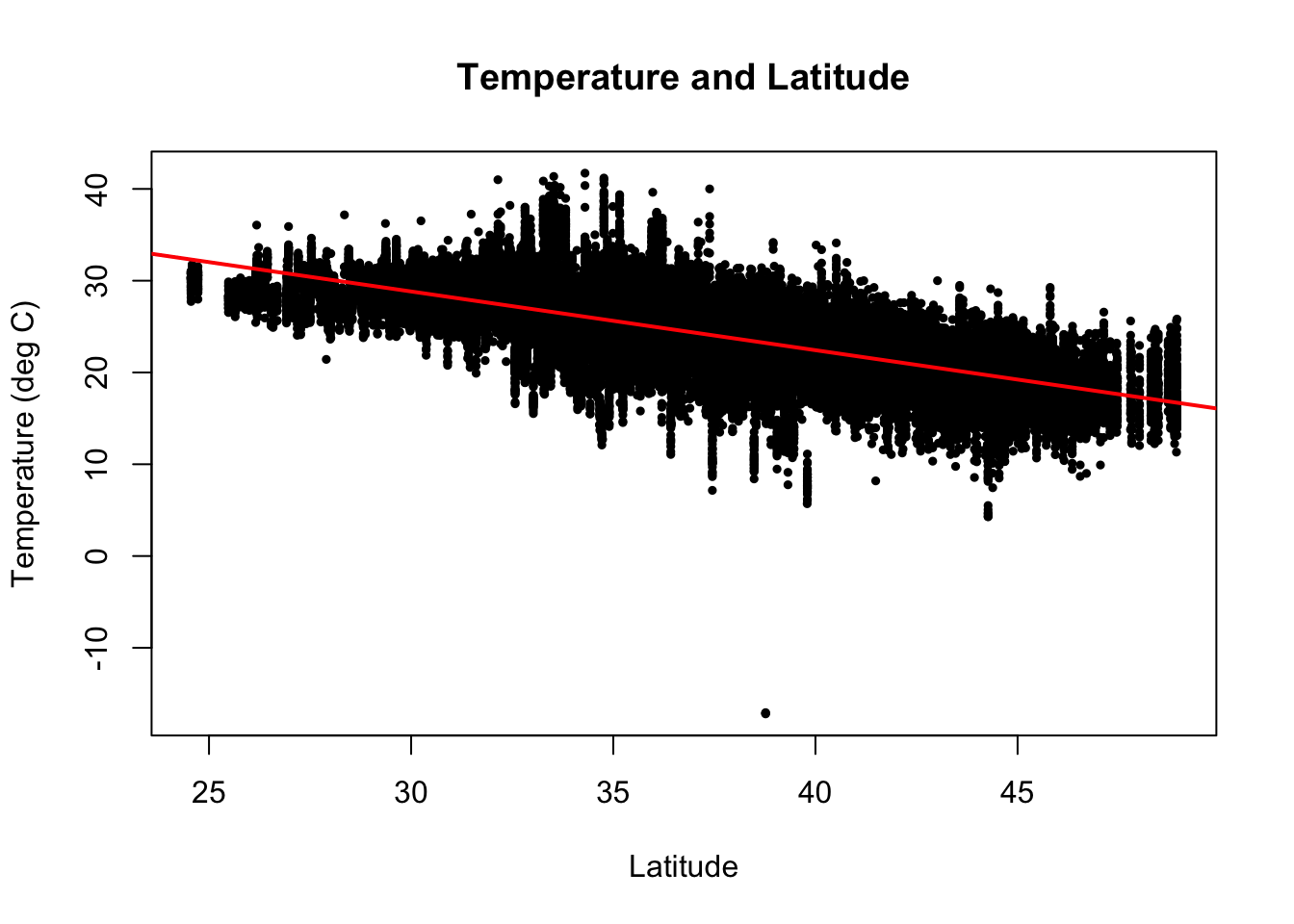
Using ggplot2 (next class)
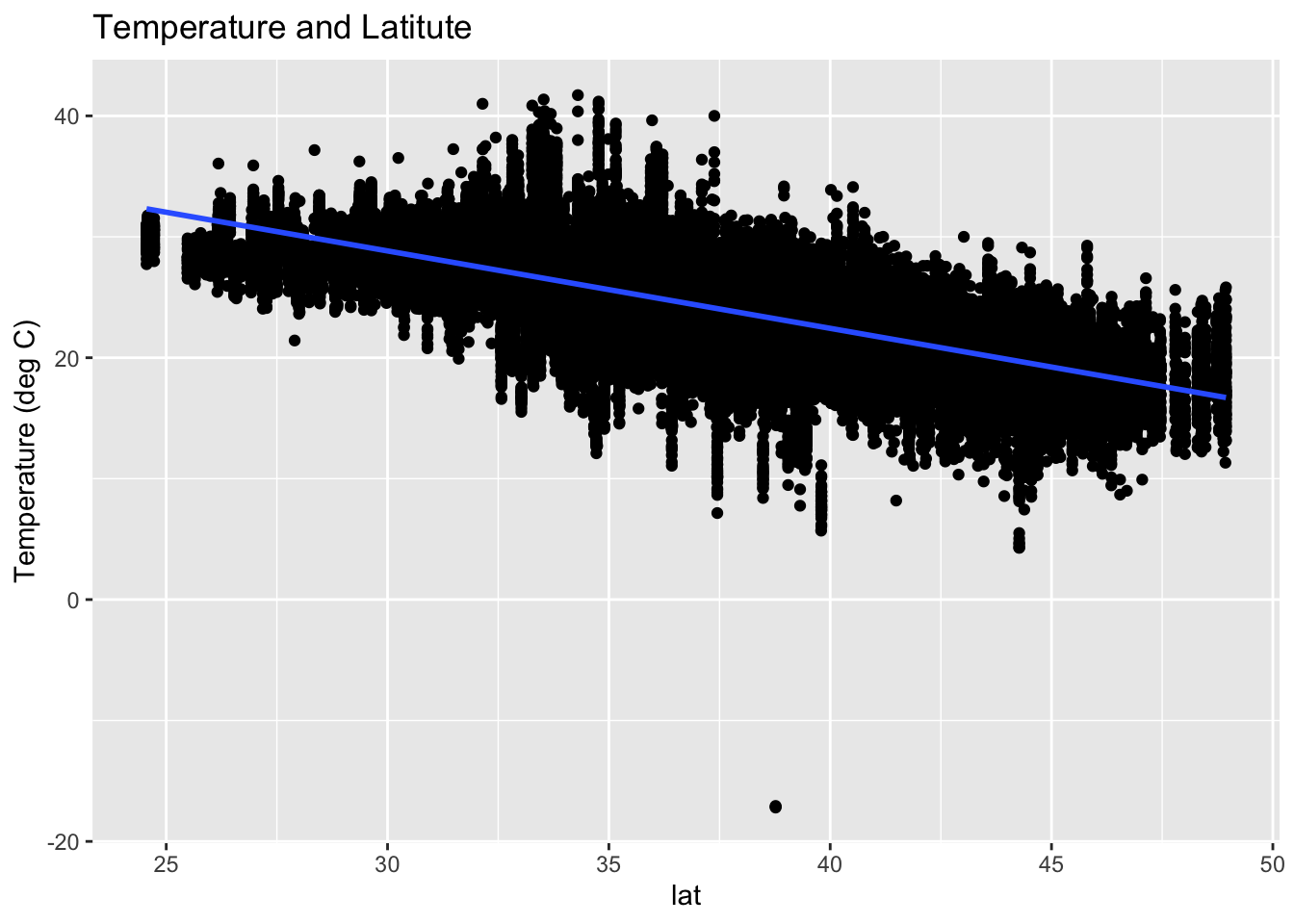
Summary
In EDA we:
- have an initial question that we aim to answer
- import, check, clean the data
- perform any data transformations to answer the initial question
- make some basic graphs to examine the data and visualize the initial question
Next topic (starting today): Tidyverse
Tidyverse
The tidyverse is not a package but a group of packages developed to work with each other.
The tidyverse makes data analysis simpler and code easier to read by sacrificing some flexibility.
One way code is simplified by ensuring all functions take and return tidy data.
Tidy data
Stored in a data frame.
Each observation is exactly one row.
Variables are stored in columns.
Not all data can be represented this way, but a very large subset of data analysis challenges are based on tidy data.
Assuming data is tidy simplifies coding and frees up our minds for statistical thinking.
Tidy data
- This is an example of a tidy dataset:
country year fertility
1 Germany 1960 2.41
2 South Korea 1960 6.16
3 Germany 1961 2.44
4 South Korea 1961 5.99
5 Germany 1962 2.47
6 South Korea 1962 5.79Tidy data
- Originally, the data was in the following format:
country 1960 1961 1962 1963 1964 1965 1966 1967 1968 1969 1970
1 Germany 2.41 2.44 2.47 2.49 2.49 2.48 2.44 2.37 2.28 2.17 2.04
2 South Korea 6.16 5.99 5.79 5.57 5.36 5.16 4.99 4.85 4.73 4.62 4.53- This is not tidy.
Tidyverse packages
tibble - improves data frame class.
readr - import data.
dplyr - used to modify data frames.
ggplot2 - simplifies plotting.
tidyr - helps convert data into tidy format.
stringr - string processing.
forcats - utilities for categorical data.
purrr - tidy version of apply functions.
dplyr
In this lecture we focus mainly on dplyr.
In particular the following functions:
mutateselectacrossfiltergroup_bysummarize
Adding a column with mutate
Notice that here we used
totalandpopulationinside the function, which are objects that are not defined in our workspace.This is known as non-standard evaluation where the context is used to know what variable names means.
Tidyverse extensively uses non-standard evaluation.
This can create confusion but it certainly simplifies code.
Subsetting with filter
Selecting columns with select
Transforming variables
The function
mutatecan also be used to transform variables.For example, the following code takes the log transformation of the population variable:
Transforming variables
Often, we need to apply the same transformation to several variables.
The function
acrossfacilitates the operation.For example if want to log transform both population and total murders we can use:
Transforming variables
The helper functions come in handy when using across.
An example is if we want to apply the same transformation to all numeric variables:
- or all character variables:
- There are several other useful helper functions.
The pipe: |> or %>%
We use the pipe to chain a series of operations.
For example if we want to select columns and then filter rows we chain like this:
\[ \mbox{original data } \rightarrow \mbox{ select } \rightarrow \mbox{ filter } \]
The pipe: |> or %>%
- The code looks like this:
state region rate
1 Hawaii West 0.5145920
2 Iowa North Central 0.6893484
3 New Hampshire Northeast 0.3798036
4 North Dakota North Central 0.5947151
5 Vermont Northeast 0.3196211The object on the left of the pipe is used as the first argument for the function on the right.
The second argument becomes the first, the third the second, and so on…
The pipe: |> or %>%
- Here is a simple example:
Summarizing data
We use the dplyr
summarizefunction, not to be confused withsummaryfrom R base.Here is an example of how it works:
- Let’s compute murder rate for the US. Is the above it?
Summarizing data
No, the rate is NOT the average of rates.
It is the total murders divided by total population:
Multiple summaries
- Suppose we want the median, minimum and max population size:
median min max
1 4339367 563626 37253956- Why don’t we use
quantiles?
Multiple summaries
Warning
Using a function that returns more than one number within summarize will soon be deprecated.
- For multiple summaries we use
reframe:
Multiple summaries
- However, if we want a column per summary, as when we called
min,median, andmaxseparately, we have to define a function that returns a data frame like this:
- Then we can call
summarize:
Group then summarize
Let’s compute murder rate by region.
Take a close look at this output?
# A tibble: 4 × 6
# Groups: region [2]
state abb region population total rate
<chr> <chr> <fct> <dbl> <dbl> <dbl>
1 Alabama AL South 4779736 135 2.82
2 Alaska AK West 710231 19 2.68
3 Arizona AZ West 6392017 232 3.63
4 Arkansas AR South 2915918 93 3.19Note the
Groups: region [4]at the top.This is a special data frame called a grouped data frame.
Group then summarize
- In particular
summarize, will behave differently when acting on this object.
# A tibble: 4 × 2
region rate
<fct> <dbl>
1 Northeast 2.66
2 South 3.63
3 North Central 2.73
4 West 2.66- The
summarizefunction applies the summarization to each group separately.
Group then summarize
- For another example, let’s compute the median, minimum, and maximum population in the four regions of the country using the
median_min_maxpreviously defined:
Group then summarize
You can also summarize a variable but not collapse the dataset.
We use
mutateinstead ofsummarize.Here is an example where we add a column with the population in each region and the number of states in the region, shown for each state.
ungroup
- When we do this, we usually want to ungroup before continuing our analysis.
- This avoids having a grouped data frame that we don’t need.
pull
- Tidyverse function always returns a data frame. Even if its just one number.
pull
- To get a numeric use pull:
Sorting data frames
- States order by rate
state abb region population total rate
1 Vermont VT Northeast 625741 2 0.3196211
2 New Hampshire NH Northeast 1316470 5 0.3798036
3 Hawaii HI West 1360301 7 0.5145920
4 North Dakota ND North Central 672591 4 0.5947151
5 Iowa IA North Central 3046355 21 0.6893484
6 Idaho ID West 1567582 12 0.7655102Sorting data frames
- If we want decreasing we can either use the negative or, for more readability, use
desc:
state abb region population total rate
1 District of Columbia DC South 601723 99 16.452753
2 Louisiana LA South 4533372 351 7.742581
3 Missouri MO North Central 5988927 321 5.359892
4 Maryland MD South 5773552 293 5.074866
5 South Carolina SC South 4625364 207 4.475323
6 Delaware DE South 897934 38 4.231937Sorting data frames
- We can use two variables as well:
state abb region population total rate
1 Pennsylvania PA Northeast 12702379 457 3.5977513
2 New Jersey NJ Northeast 8791894 246 2.7980319
3 Connecticut CT Northeast 3574097 97 2.7139722
4 New York NY Northeast 19378102 517 2.6679599
5 Massachusetts MA Northeast 6547629 118 1.8021791
6 Rhode Island RI Northeast 1052567 16 1.5200933
7 Maine ME Northeast 1328361 11 0.8280881
8 New Hampshire NH Northeast 1316470 5 0.3798036
9 Vermont VT Northeast 625741 2 0.3196211
10 District of Columbia DC South 601723 99 16.4527532
11 Louisiana LA South 4533372 351 7.7425810